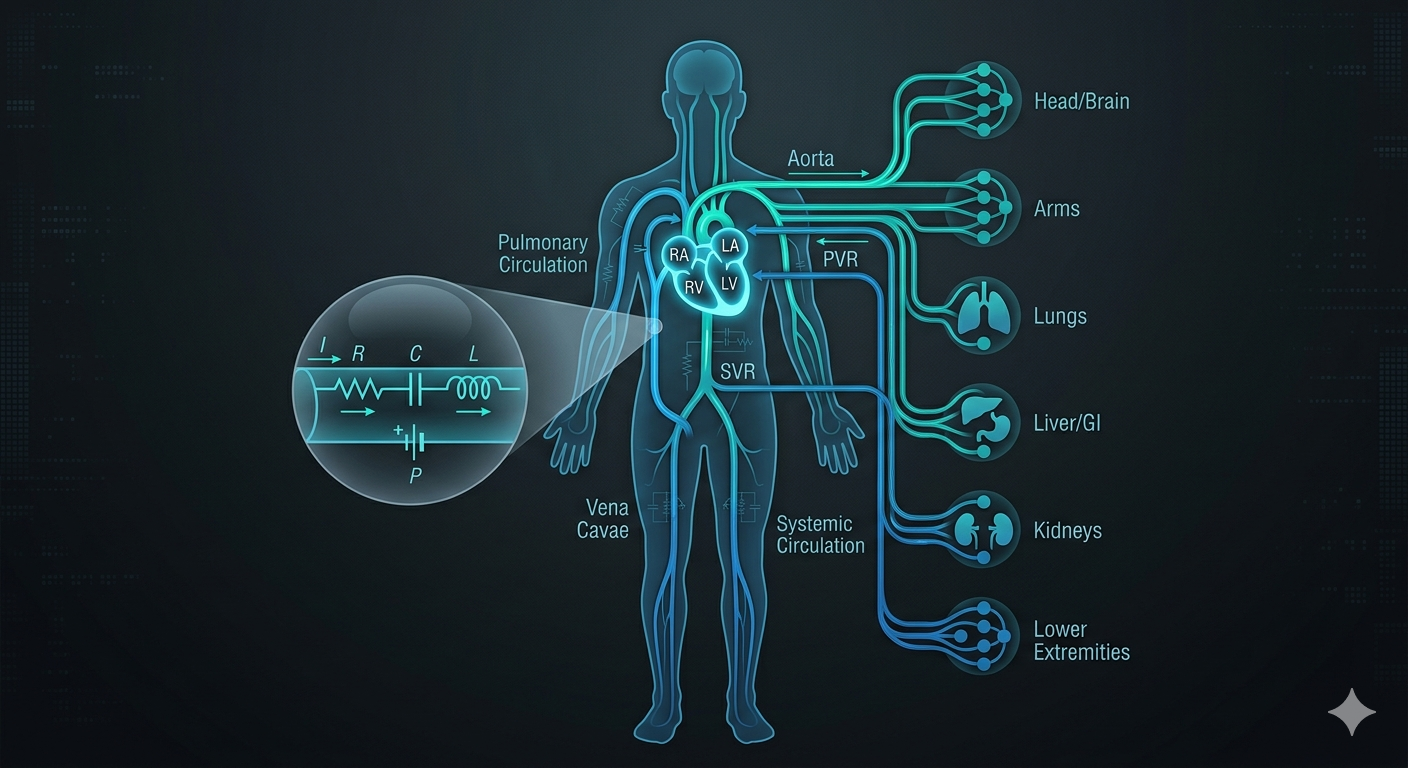
Whole-body cardiovascular digital twin
Closed-loop, lumped-parameter (RCR) network for whole-body hemodynamics—pressure, flow, and disease-state response.
Postdoctoral Research Associate
I build physiologically-grounded digital twins of human health and disease—predictive models that translate complex physiology into clinical insight and de-risk drug development. In parallel, I develop AI agents that accelerate how these models are constructed, calibrated, and refined.

Research
Computational frameworks bridging physiology, pharmacokinetics, and AI-assisted model authoring.

Closed-loop, lumped-parameter (RCR) network for whole-body hemodynamics—pressure, flow, and disease-state response.
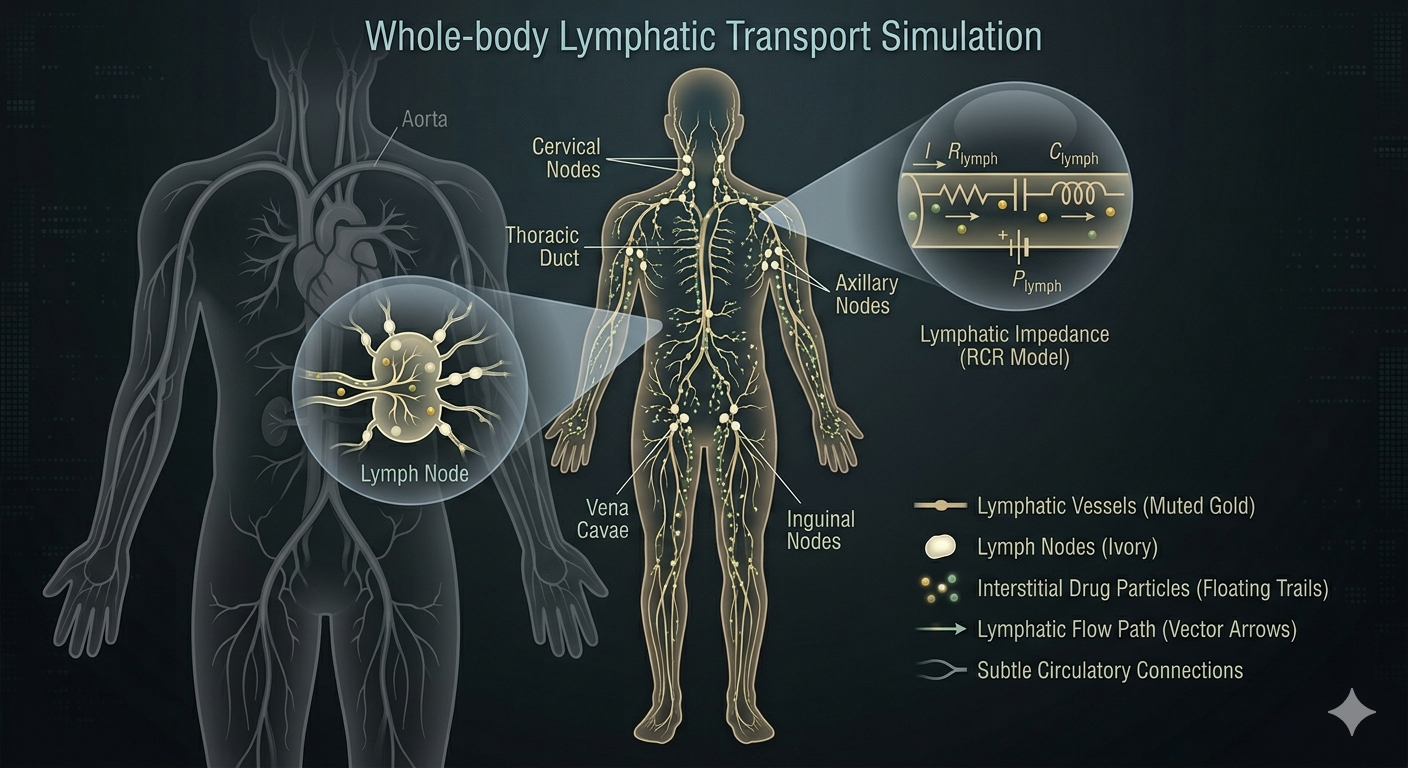
Comprehensive framework for interstitial and lymphatic drug transport—essential for subcutaneous biologics and immuno-oncology PK.
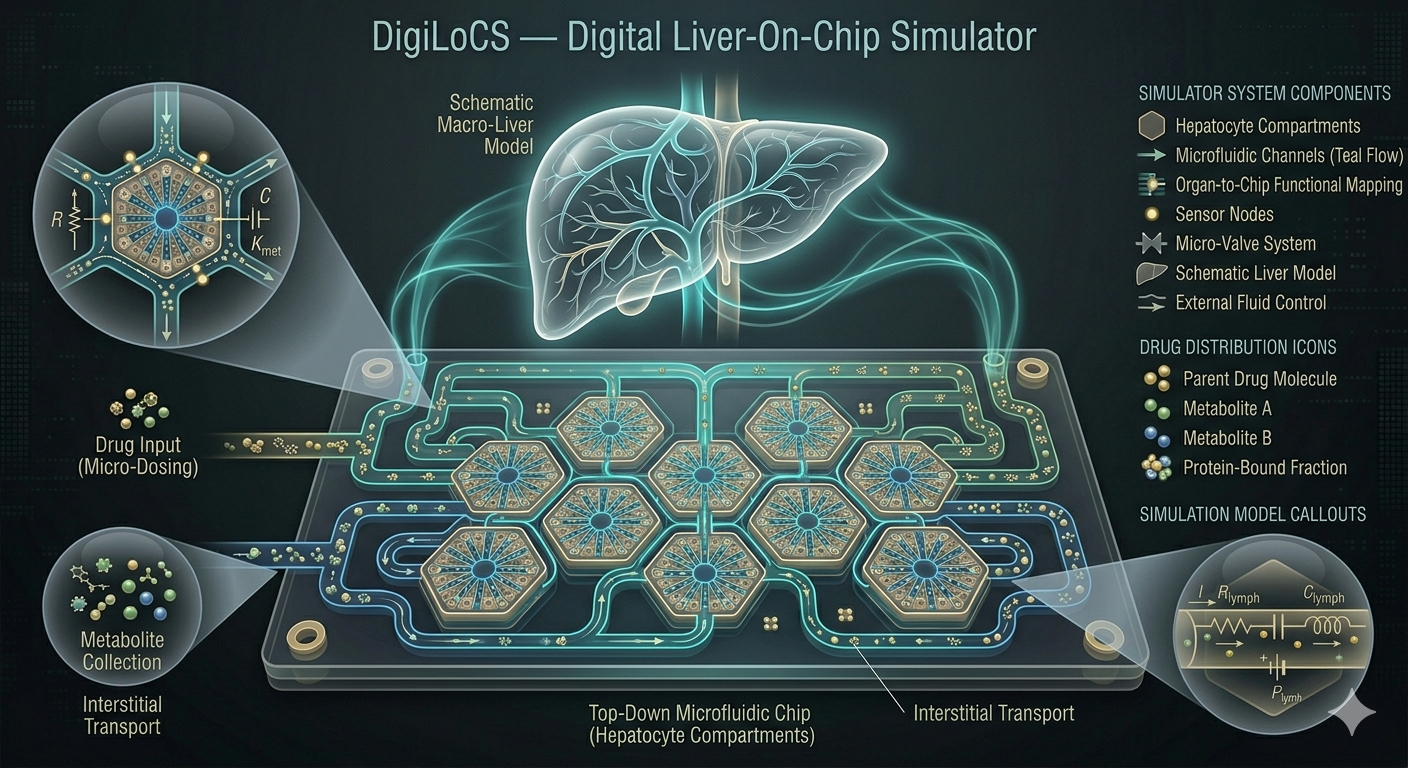
Predictive organ-on-chip simulator differentiating active vs. passive hepatic clearance for IVIVE. Published in PLOS ONE.
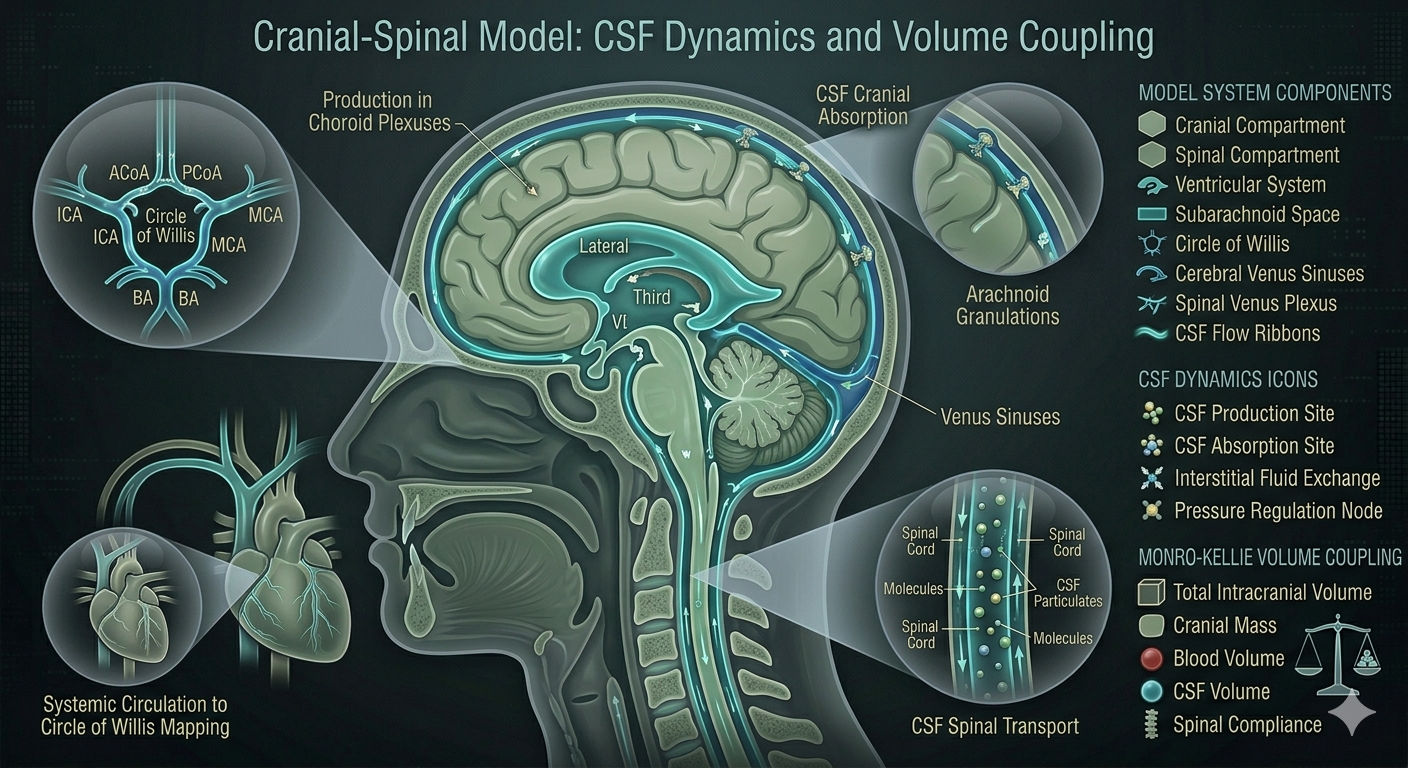
Coupled lumped-parameter model capturing Monro–Kellie volume coupling, intracranial pulsatility, and cerebrovascular regulation—relevant to intrathecal CNS delivery.
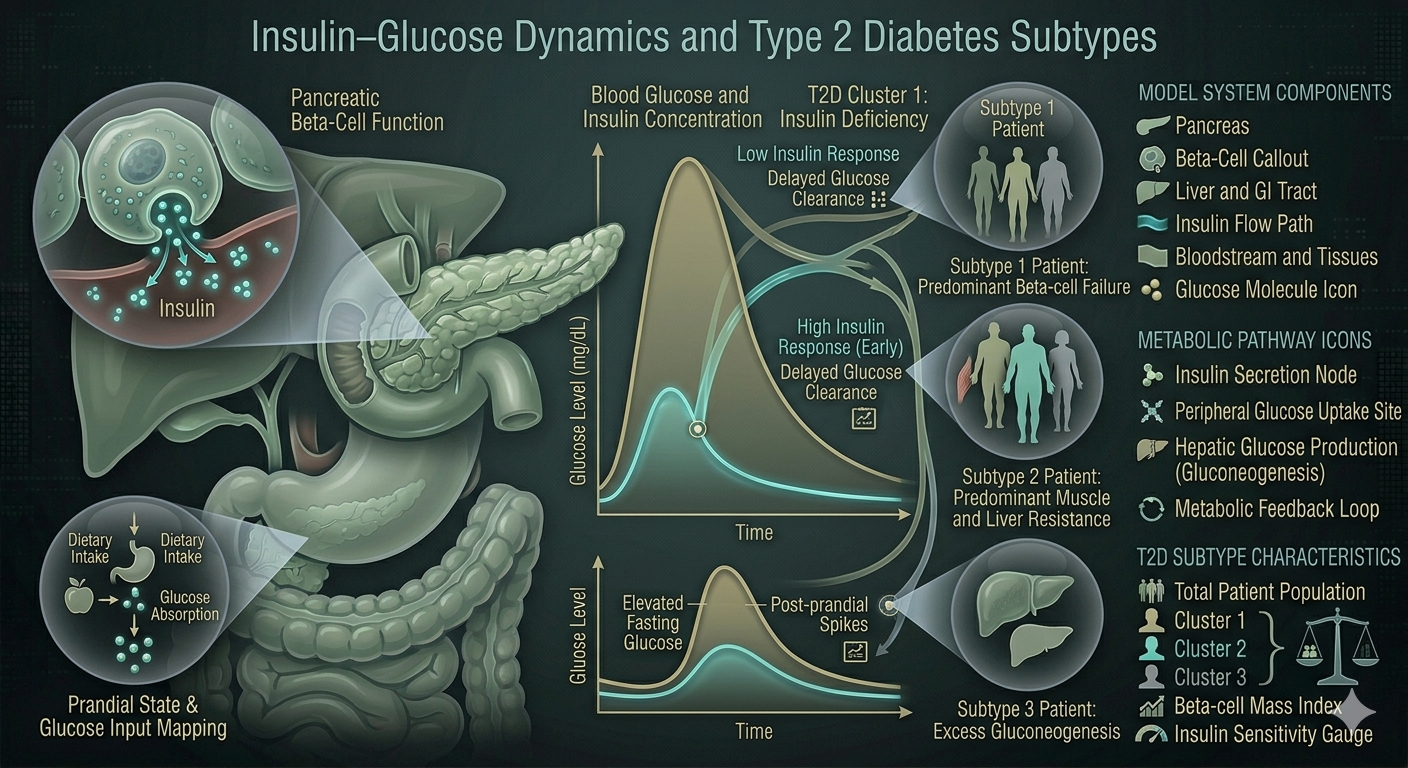
Extended oral minimal model with adipokine (leptin) and BMI coupling; identified three patho-clinical clusters in uncontrolled T2D patients.
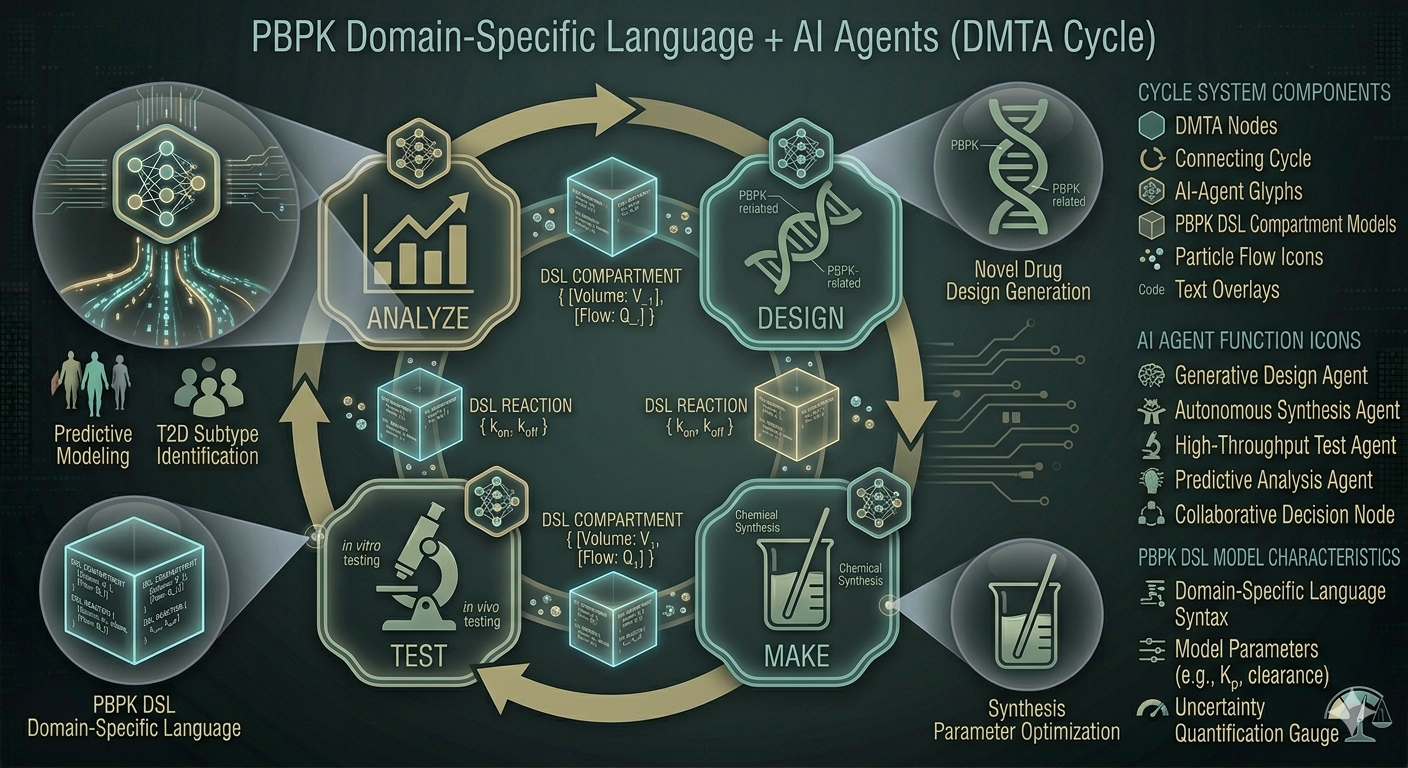
A domain-specific language for compartmental PK models that integrates a hierarchy of AI agents into the Design–Make–Test–Analyze (DMTA) cycle.